
Sandra Camelo-Piragua, MD, Assistant Professor of Neuropathology, joined the Department of Pathology in 2010. Her clinical practice focuses on diagnosing diseases of the peripheral and central nervous systems and muscle, in both adult and pediatric patients.
Having studied on three continents, Camelo-Piragua now enjoys teaching and is the director of the neuropathology fellowship. She Is interested in developing information technology tools that increase access to neuropathology education.
Working in close collaboration, Camelo-Piragua and assistant professor of neurosurgery, Daniel A. Orringer, MD, along with their colleagues, have developed a new technique for diagnosing brain tumors.
The technology uses stimulated Raman scattering (SRS) microscopy to create Simulated Raman Histology (SRH). In SRH, fresh tissue is collected during surgery, placed on glass slides without further processing or staining and scoped using a fiber-laser-based SRS microscopy. The acquired images are virtually colored to highlight the features of brain tumors. This results in images that very closely resemble traditional hematoxylin and eosin (H&E) stained tissue but reduces the process from 30-minutes to about 3.
In a study published online in early February in Nature Biomedical Engineering, expert neuropathologists from different institutions were given images obtained at the time of surgery and processed using either traditional H&E staining or SRH methods. There was a near-perfect concordance between the H&E method of diagnosis and SRH, which was 92% accurate.
Originally studied for its potential capability to guide surgeons to identify microscopic tumor tissue during surgery, this groundbreaking technique can be used to determine types of tumors as well.
Camelo-Piragua equates pathologists to detectives trying to solve a puzzle. She explains, “SRH imaging will ensure that appropriate and good quality tissue is collected to reach our ultimate goal: accurate diagnoses.”
Orringer and his students are pushing the boundaries of what this technique offers. They are using machine learning algorithms to teach a computer to recognize different types of tumors. “The more we feed the computer, the more accurate the diagnoses will become,” he says.
There are approximately 1,400 institutions performing brain surgery in the U.S. but fewer than 800 board-certified neuropathologists. Use of SRH could potentially expand the capabilities of hospitals without experts on-site because the images could be reviewed remotely.
While the current prototype is for research only, a large-scale clinical trial could be the next step. “The future is bright,” says Camelo-Piragua “and I am glad I am contributing in moving forward and seeing things with new, fresh, and different eyes.”
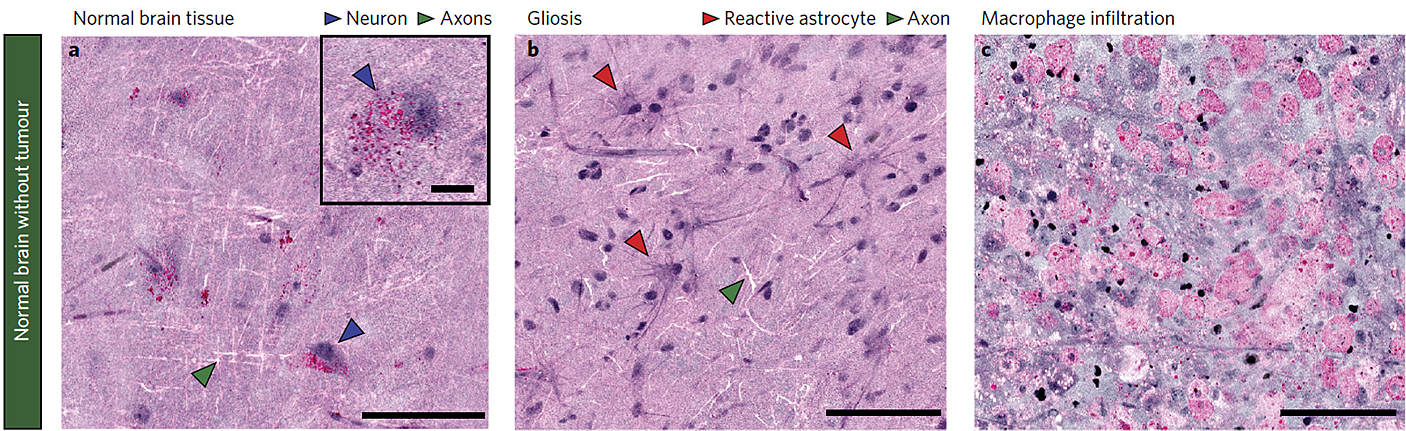 Figure from Nature Biomedical Engineer 2017 DOI: 10.1038/s41551-016-0027
Figure from Nature Biomedical Engineer 2017 DOI: 10.1038/s41551-016-0027
As a young child, my father came home with a book about how the brain works. It was authored by a Colombian physician, Dr. Rodolfo Llinas, working at NYU and NASA. He became a role model, and his book opened my eyes to the wonders of neuroscience. As a medical student, I became involved with Alzheimer’s disease-related research and later had the opportunity to come to Harvard Medical school and spend a couple of years studying different neurodegenerative diseases. When deciding what residency to pursue, I looked for one that would allow me time to study and analyze the most complex and unusual cases.
I have always loved understanding how things work. During my research years, after medical school, I realized I enjoyed understanding histological patterns under the microscope. The microscope became a new window, a different type of book for learning and understanding those things that were hidden in our bodies. Neuropathology became a logical passion to follow.
As a physician, my ultimate goal is to serve the patient. My role as a neuropathologist is to use all my knowledge and expertise to provide the most accurate diagnosis. This will translate into appropriate therapeutic measurements in that individual. Working at the University of Michigan I have had the opportunity to fulfill my desire to serve this clinical purpose. Furthermore, I had the opportunity to interact and collaborate with colleagues within and outside of my department and have had the chance to try new and bold ideas.
I love the open collaborative spirit of the faculty at the University of Michigan. Having the opportunity to work with colleagues outside your area of expertise allows you to grow intellectually and personally. There is always something new to learn. Collaborating outside your silo allows you to see the world with different eyes; helps you tackle the problems from different perspectives and opens your mind to new possibilities.
Walking or biking on the trails of the numerous parks and natural spots in Ann Arbor with my family. Camping with my family and discovering Michigan beauties.
The public libraries and bookstores. I could spend an entire day there. I like to browse, learn, and discover new things outside pathology.
—
Additional authors from Michigan Medicine include Balaji Pandian, Yashar S. Niknafs, Todd C. Hollon, Julianne Boyle, Spencer Lewis, Mia Garrard, Shawn L. Hervey-Jumper, Hugh J. L. Garton, Cormac O. Maher, Jason A. Heth, Oren Sagher, D. Andrew Wilkinson, Sriram Venneti, Kathryn A. McFadden, Amanda Fisher-Hubbard, Andrew P. Lieberman and Timothy D. Johnson. Matija Snuderl is from New York University. Shakti H. Ramkissoon is from Harvard Medical School and the Dana Farber Cancer Institute. X. Sunney Xie is from Harvard University. Jay K. Trautman and Christian W. Freudiger are from Invenio Imaging.
Disclosure: Orringer and co-author Xie are advisers and shareholders of Invenio Imaging Inc., a company developing SRS microscopy systems. Co-author Freudiger is an employee and shareholder of Invenio Imaging, Inc.
The University of Michigan, Harvard University, and Invenio Imaging have intellectual property used in this research.
 ON THE COVER
ON THE COVER
Breast team reviewing a patient's slide. (From left to right) Ghassan Allo, Fellow; Laura Walters, Clinical Lecturer; Celina Kleer, Professor. See Article 2014Department Chair |

newsletter
INSIDE PATHOLOGYAbout Our NewsletterInside Pathology is an newsletter published by the Chairman's Office to bring news and updates from inside the department's research and to become familiar with those leading it. It is our hope that those who read it will enjoy hearing about those new and familiar, and perhaps help in furthering our research. CONTENTS
|
 ON THE COVER
ON THE COVER
Autopsy Technician draws blood while working in the Wayne County morgue. See Article 2016Department Chair |

newsletter
INSIDE PATHOLOGYAbout Our NewsletterInside Pathology is an newsletter published by the Chairman's Office to bring news and updates from inside the department's research and to become familiar with those leading it. It is our hope that those who read it will enjoy hearing about those new and familiar, and perhaps help in furthering our research. CONTENTS
|
 ON THE COVER
ON THE COVER
Dr. Sriram Venneti, MD, PhD and Postdoctoral Fellow, Chan Chung, PhD investigate pediatric brain cancer. See Article 2017Department Chair |

newsletter
INSIDE PATHOLOGYAbout Our NewsletterInside Pathology is an newsletter published by the Chairman's Office to bring news and updates from inside the department's research and to become familiar with those leading it. It is our hope that those who read it will enjoy hearing about those new and familiar, and perhaps help in furthering our research. CONTENTS
|
 ON THE COVER
ON THE COVER
Director of the Neuropathology Fellowship, Dr. Sandra Camelo-Piragua serves on the Patient and Family Advisory Council. 2018Department Chair |

newsletter
INSIDE PATHOLOGYAbout Our NewsletterInside Pathology is an newsletter published by the Chairman's Office to bring news and updates from inside the department's research and to become familiar with those leading it. It is our hope that those who read it will enjoy hearing about those new and familiar, and perhaps help in furthering our research. CONTENTS
|
 ON THE COVER
ON THE COVER
Residents Ashley Bradt (left) and William Perry work at a multi-headed scope in our new facility. 2019Department Chair |

newsletter
INSIDE PATHOLOGYAbout Our NewsletterInside Pathology is an newsletter published by the Chairman's Office to bring news and updates from inside the department's research and to become familiar with those leading it. It is our hope that those who read it will enjoy hearing about those new and familiar, and perhaps help in furthering our research. CONTENTS
|
 ON THE COVER
ON THE COVER
Dr. Kristine Konopka (right) instructing residents while using a multi-headed microscope. 2020Department Chair |

newsletter
INSIDE PATHOLOGYAbout Our NewsletterInside Pathology is an newsletter published by the Chairman's Office to bring news and updates from inside the department's research and to become familiar with those leading it. It is our hope that those who read it will enjoy hearing about those new and familiar, and perhaps help in furthering our research. CONTENTS
|
 ON THE COVER
ON THE COVER
Patient specimens poised for COVID-19 PCR testing. 2021Department Chair |

newsletter
INSIDE PATHOLOGYAbout Our NewsletterInside Pathology is an newsletter published by the Chairman's Office to bring news and updates from inside the department's research and to become familiar with those leading it. It is our hope that those who read it will enjoy hearing about those new and familiar, and perhaps help in furthering our research. CONTENTS
|
 ON THE COVER
ON THE COVER
Dr. Pantanowitz demonstrates using machine learning in analyzing slides. 2022Department Chair |

newsletter
INSIDE PATHOLOGYAbout Our NewsletterInside Pathology is an newsletter published by the Chairman's Office to bring news and updates from inside the department's research and to become familiar with those leading it. It is our hope that those who read it will enjoy hearing about those new and familiar, and perhaps help in furthering our research. CONTENTS
|
 ON THE COVER
ON THE COVER
(Left to Right) Drs. Angela Wu, Laura Lamps, and Maria Westerhoff. 2023Department Chair |

newsletter
INSIDE PATHOLOGYAbout Our NewsletterInside Pathology is an newsletter published by the Chairman's Office to bring news and updates from inside the department's research and to become familiar with those leading it. It is our hope that those who read it will enjoy hearing about those new and familiar, and perhaps help in furthering our research. CONTENTS
|
 ON THE COVER
ON THE COVER
Illustration representing the various machines and processing used within our labs. 2024Department Chair |

newsletter
INSIDE PATHOLOGYAbout Our NewsletterInside Pathology is an newsletter published by the Chairman's Office to bring news and updates from inside the department's research and to become familiar with those leading it. It is our hope that those who read it will enjoy hearing about those new and familiar, and perhaps help in furthering our research. CONTENTS
|
 ON THE COVER
ON THE COVER
Rendering of the D. Dan and Betty Khn Health Care Pavilion. Credit: HOK 2025Department Chair |

newsletter
INSIDE PATHOLOGYAbout Our NewsletterInside Pathology is an newsletter published by the Chairman's Office to bring news and updates from inside the department's research and to become familiar with those leading it. It is our hope that those who read it will enjoy hearing about those new and familiar, and perhaps help in furthering our research. CONTENTS
|

MLabs, established in 1985, functions as a portal to provide pathologists, hospitals. and other reference laboratories access to the faculty, staff and laboratories of the University of Michigan Health System’s Department of Pathology. MLabs is a recognized leader for advanced molecular diagnostic testing, helpful consultants and exceptional customer service.