

By Kara Gavin | www.uofmhealth.org | 13 September
ANN ARBOR, Mich. — If you get pneumonia, or even an infected cut, your body is now a war zone.
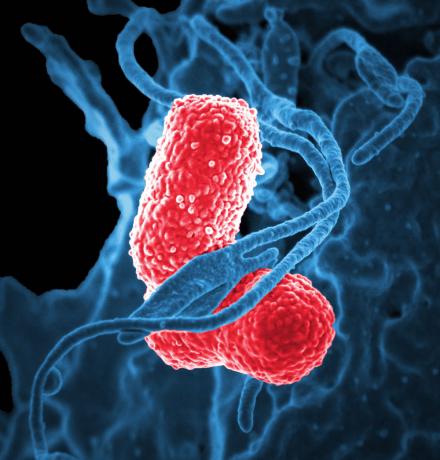
And as your immune system battles the invading bacteria, the outcome of that war may hinge on a microscopic arms race based not on missiles or bombs, but on an essential element: iron.
Now, scientists from the University of Michigan Medical School say they have figured out how that race for iron actually increases the risk we face from one of our most dangerous microscopic foes, Klebsiella pneumoniae. They made the discovery in mice with pneumonia.
K. pneumoniae has already figured out how to overcome our best defenses – including, in some cases, all our most powerful antibiotics.
It’s the third-most-common cause of infections that arise in hospitalized patients, and causes pneumonia, urinary tract infections, wound infections and bloodstream infections. In its most drug-resistant form, it’s considered one of the carbapenem resistant Enterobacteriaceae, or CRE – so called "nightmare bacteria".
The new findings could aid the search for drugs to fight it, and other "superbugs".
In a new paper in the journal mBio, the team led by Michael Bachman, M.D., Ph.D. report what happens after the bacteria send out tiny iron-scavenging molecules.
Called siderophores, those molecules have long been thought of as a way for bacteria to gather a precious element needed to grow and reproduce, by attaching to iron and stealing it from us.
Siderophores from K. pneumoniae are hundreds of times more powerful at grabbing iron than the proteins that our own bodies produce. What’s more, the bacteria produce a kind of siderophore that our defense systems can’t neutralize.
But Bachman and his colleagues show that bacterial siderophores do much more than just grab iron. Their experiments show that K. pneumoniae actually use the molecules to help them invade the rest of the body beyond their initial point of entry, and to bring on inflammation caused by our own immune system.

"This is a bacterium that has evolved new ways to get iron, and it turns out that the mechanism it uses also causes cellular stress during infections," says Bachman, an assistant professor of pathology at U-M. "That response triggers an immune response that tells our bodies to fight the infection, but it also activates a mechanism that allows bacteria to escape and travel to the rest of the body."
That mechanism, a protein called Hif-1alpha, normally helps our bodies respond to low oxygen or low iron. But when K. pneumoniae siderophores activate it, it worsens the infection. Exactly how is still a mystery.
Bachman sees the effects of K. pneumoniae on patients in his work as a clinical microbiologist at the U-M Health System, where he’s associate director of clinical microbiology. That motivates him to study it in the lab, though he notes that it’s too early to say exactly how the discovery could be used to help patients.
One promising route could be to develop strategies to prevent the bacteria from sending out siderophores in the first place -- or to use the siderophores to bring antibiotics back into bacterial cells. Or, it may even be possible to create a vaccine based on siderophores to teach the immune system to attack them as invaders.
Meanwhile, he and others at U-M including Microbiology & Immunology chair Harry Mobley, Ph.D., are pursuing a broader goal through U-M’s Host Microbiome Initiative. Using an advanced genetic technique called transposon sequencing, they’re working to figure out which genes K. pneumoniae and other superbugs absolutely need in order to cause infections.
To figure out the double role that siderophores play in a K. pneumoniae infection, Bachman and his colleagues had to figure out how to create bacteria that would not reproduce, while still producing siderophores. They were able to mutate bacteria in a way that kept iron-laden siderophores from getting back into the bacterial cells and promoting growth. That way, the number of bacteria in a mouse’s body stayed the same, but the researchers could study what the siderophores did.
They expected that if the siderophores grabbed enough iron from the mouse’s lung tissue and robbed the mouse an essential element, the mouse’s cells would respond.. Indeed, they found that the mouse immune system was triggered directly by the siderophores, causing the release of molecules called cytokines that attract infection-fighting cells and cause inflammation as a result. The activation of Hif-1alpha was part of this – because the mouse cells sensed that iron supply was low.
So, the team figured out how to eliminate Hif-1alpha in the cells lining the mouse lungs – and showed that in these mice, most of the bacteria couldn’t escape from the lungs to the spleen. But in mice that made Hif-1alpha, more K. pneumoniae bacteria spread to the rest of the body, making the infection worse.
"This work provides us with motivation to target the production of siderophores, rather than just the uptake of them," says Bachman. "Now that we know that the bacteria cause cellular stress just by secreting them, we may be able to prevent these effects if we neutralize them."
The research was funded by the Natural Sciences and Engineering Research Council of Canada, through a grant to co-author Martin Dozois. Dozois and co-author Sebastien Houle are at the Institut national de la recherche scientifique, Institut Armand-Frappier, in Laval, Quebec. Recent U-M Ph.D. recipient Victoria Holden is the study’s first author, and the authors also include former research technician Paul Breen. Reference: mBio, 7(6):e01397-16, doi:10.1128/mBio.01397-16.
 ON THE COVER
ON THE COVER
Breast team reviewing a patient's slide. (From left to right) Ghassan Allo, Fellow; Laura Walters, Clinical Lecturer; Celina Kleer, Professor. See Article 2014Department Chair |

newsletter
INSIDE PATHOLOGYAbout Our NewsletterInside Pathology is an newsletter published by the Chairman's Office to bring news and updates from inside the department's research and to become familiar with those leading it. It is our hope that those who read it will enjoy hearing about those new and familiar, and perhaps help in furthering our research. CONTENTS
|
 ON THE COVER
ON THE COVER
Autopsy Technician draws blood while working in the Wayne County morgue. See Article 2016Department Chair |

newsletter
INSIDE PATHOLOGYAbout Our NewsletterInside Pathology is an newsletter published by the Chairman's Office to bring news and updates from inside the department's research and to become familiar with those leading it. It is our hope that those who read it will enjoy hearing about those new and familiar, and perhaps help in furthering our research. CONTENTS
|
 ON THE COVER
ON THE COVER
Dr. Sriram Venneti, MD, PhD and Postdoctoral Fellow, Chan Chung, PhD investigate pediatric brain cancer. See Article 2017Department Chair |

newsletter
INSIDE PATHOLOGYAbout Our NewsletterInside Pathology is an newsletter published by the Chairman's Office to bring news and updates from inside the department's research and to become familiar with those leading it. It is our hope that those who read it will enjoy hearing about those new and familiar, and perhaps help in furthering our research. CONTENTS
|
 ON THE COVER
ON THE COVER
Director of the Neuropathology Fellowship, Dr. Sandra Camelo-Piragua serves on the Patient and Family Advisory Council. 2018Department Chair |

newsletter
INSIDE PATHOLOGYAbout Our NewsletterInside Pathology is an newsletter published by the Chairman's Office to bring news and updates from inside the department's research and to become familiar with those leading it. It is our hope that those who read it will enjoy hearing about those new and familiar, and perhaps help in furthering our research. CONTENTS
|
 ON THE COVER
ON THE COVER
Residents Ashley Bradt (left) and William Perry work at a multi-headed scope in our new facility. 2019Department Chair |

newsletter
INSIDE PATHOLOGYAbout Our NewsletterInside Pathology is an newsletter published by the Chairman's Office to bring news and updates from inside the department's research and to become familiar with those leading it. It is our hope that those who read it will enjoy hearing about those new and familiar, and perhaps help in furthering our research. CONTENTS
|
 ON THE COVER
ON THE COVER
Dr. Kristine Konopka (right) instructing residents while using a multi-headed microscope. 2020Department Chair |

newsletter
INSIDE PATHOLOGYAbout Our NewsletterInside Pathology is an newsletter published by the Chairman's Office to bring news and updates from inside the department's research and to become familiar with those leading it. It is our hope that those who read it will enjoy hearing about those new and familiar, and perhaps help in furthering our research. CONTENTS
|
 ON THE COVER
ON THE COVER
Patient specimens poised for COVID-19 PCR testing. 2021Department Chair |

newsletter
INSIDE PATHOLOGYAbout Our NewsletterInside Pathology is an newsletter published by the Chairman's Office to bring news and updates from inside the department's research and to become familiar with those leading it. It is our hope that those who read it will enjoy hearing about those new and familiar, and perhaps help in furthering our research. CONTENTS
|
 ON THE COVER
ON THE COVER
Dr. Pantanowitz demonstrates using machine learning in analyzing slides. 2022Department Chair |

newsletter
INSIDE PATHOLOGYAbout Our NewsletterInside Pathology is an newsletter published by the Chairman's Office to bring news and updates from inside the department's research and to become familiar with those leading it. It is our hope that those who read it will enjoy hearing about those new and familiar, and perhaps help in furthering our research. CONTENTS
|
 ON THE COVER
ON THE COVER
(Left to Right) Drs. Angela Wu, Laura Lamps, and Maria Westerhoff. 2023Department Chair |

newsletter
INSIDE PATHOLOGYAbout Our NewsletterInside Pathology is an newsletter published by the Chairman's Office to bring news and updates from inside the department's research and to become familiar with those leading it. It is our hope that those who read it will enjoy hearing about those new and familiar, and perhaps help in furthering our research. CONTENTS
|
 ON THE COVER
ON THE COVER
Illustration representing the various machines and processing used within our labs. 2024Department Chair |

newsletter
INSIDE PATHOLOGYAbout Our NewsletterInside Pathology is an newsletter published by the Chairman's Office to bring news and updates from inside the department's research and to become familiar with those leading it. It is our hope that those who read it will enjoy hearing about those new and familiar, and perhaps help in furthering our research. CONTENTS
|
 ON THE COVER
ON THE COVER
Rendering of the D. Dan and Betty Khn Health Care Pavilion. Credit: HOK 2025Department Chair |

newsletter
INSIDE PATHOLOGYAbout Our NewsletterInside Pathology is an newsletter published by the Chairman's Office to bring news and updates from inside the department's research and to become familiar with those leading it. It is our hope that those who read it will enjoy hearing about those new and familiar, and perhaps help in furthering our research. CONTENTS
|

MLabs, established in 1985, functions as a portal to provide pathologists, hospitals. and other reference laboratories access to the faculty, staff and laboratories of the University of Michigan Health System’s Department of Pathology. MLabs is a recognized leader for advanced molecular diagnostic testing, helpful consultants and exceptional customer service.